Classes |
Public Types |
Public Member Functions |
Static Protected Member Functions |
Friends |
List of all members
npstat::AbsClassicalOrthoPoly1D Class Referenceabstract
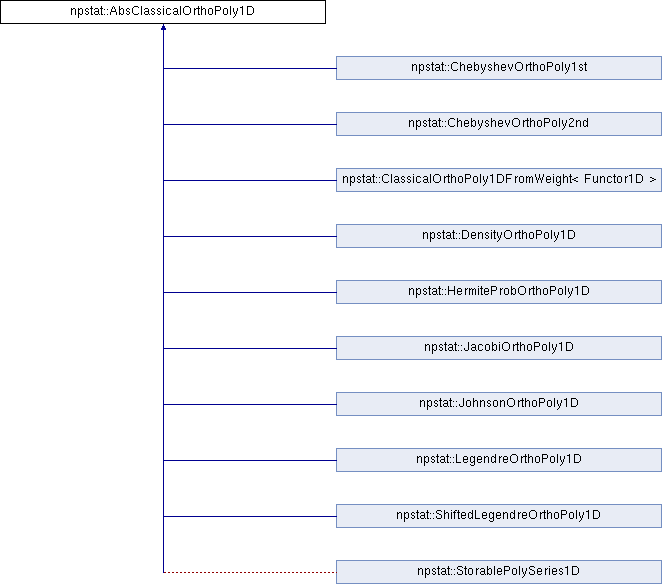
Inheritance diagram for npstat::AbsClassicalOrthoPoly1D:
Member Function Documentation◆ allmonic()
Values of all monic orthonormal polynomials up to some degree. The size of the "values" array should be at least maxdeg + 1. ◆ allpoly()
Values of all orthonormal polynomials up to some degree. Faster than calling "poly" multiple times. The size of the "values" array should be at least maxdeg + 1. ◆ calculateCoeffs()
template<class Functor , class Quadrature >
Build the coefficients of the orthogonal polynomial series for the given function. The length of the array "coeffs" should be at least maxdeg + 1. Note that the coefficients are returned in the order of increasing degree (same order is used by the "series" function). ◆ clone()
Virtual copy constructor Implemented in npstat::JohnsonOrthoPoly1D, npstat::DensityOrthoPoly1D, npstat::HermiteProbOrthoPoly1D, npstat::ChebyshevOrthoPoly2nd, npstat::ChebyshevOrthoPoly1st, npstat::JacobiOrthoPoly1D, npstat::ShiftedLegendreOrthoPoly1D, npstat::LegendreOrthoPoly1D, and npstat::ClassicalOrthoPoly1DFromWeight< Functor1D >. ◆ directQuadrature() [1/2]
template<class Quadrature >
Direct unweighted integration of of a product of multiple orthonormal poly values. The integration function should support the "integrate" method without limits. Intended for use with GaussHermiteQuadrature and such. "checkedForZeros" argument should be set to "true" if we are sure that there are no zero elements in the "degrees" array. ◆ directQuadrature() [2/2]
template<class Quadrature >
Direct unweighted integration of of a product of four orthonormal poly values. The integration function should support the "integrate" method without limits. Intended for use with GaussHermiteQuadrature and such. ◆ empiricalKroneckerDelta()
template<class Quadrature >
This method is useful for testing the numerical precision of the orthonormalization procedure. It returns the scalar products between various polynomials. ◆ empiricalKroneckerMatrix()
A faster way to generate Kronecker deltas if the whole matrix of them is needed. The "maxdeg" argument must not exceed the return value of the "maxDegree" function. The resulting matrix will be dimensioned (maxdeg + 1) x (maxdeg + 1). Note that this function will produce meaningful results only for polynomials supported on compact intervals. ◆ extWeightAverages()
template<class Functor , typename Real >
Averages of the polynomial values weighted by an external weight function. The length of array "averages" should be at least maxdeg + 1. ◆ extWeightProductAverages()
template<class Functor >
Pairwise products of the polynomial values weighted by an external weight function. The returned matrix will be symmetric and will have the dimensions (maxdeg + 1) x (maxdeg + 1). ◆ integratePoly()
An unweighted integral of the orthonormal polynomial with the given degree to some power. For this method, it is assumed that the polynomials are supported on a closed interval (without such an assumption unweighted integrals do not make much sense) and that Gauss-Legendre quadratures can be used. ◆ integrateSeries()
Integral of the series from xmin() to xmax() ◆ jacobiMatrix()
Generate principal minor of order n of the Jacobi matrix. n must not exceed the output of "maxDegree()" method. ◆ jointAverage() [1/2]
template<class Quadrature >
A measure-weighted average of a product of multiple orthonormal poly values. "checkedForZeros" argument should be set to "true" if we are sure that there are no zero elements in the "degrees" array. ◆ jointAverage() [2/2]
template<class Quadrature >
A measure-weighted average of a product of four orthonormal poly values ◆ jointIntegral()
An unweighted integral of a product of multiple orthonormal polynomials. For this method, it is assumed that the polynomials are supported on a closed interval (without such an assumption unweighted integrals do not make much sense) and that Gauss-Legendre quadratures can be used. ◆ jointSampleAverage()
template<class Numeric >
Unweighted average of a product of polynomial values for the given sample. "degrees" is the collection of polynomial degrees. Polynomials of these degrees will be included in the product. ◆ maxDegree()
Maximum polynomial degree supported Reimplemented in npstat::JohnsonOrthoPoly1D, npstat::DensityOrthoPoly1D, npstat::StorablePolySeries1D, and npstat::ClassicalOrthoPoly1DFromWeight< Functor1D >. ◆ monic()
Values of the corresponding monic polynomials ◆ poly()
Polynomial values ◆ sampleAverages()
template<typename Numeric , typename Real >
Unweighted averages of the polynomial values for the given sample. The length of array "averages" should be at least maxdeg + 1. ◆ sampleCoeffCovariance() [1/2]
template<class Numeric >
Estimate the covariances of the coefficients returned by the "sampleCoeffs" function. The "coeffs" array should be at least maxdeg + 1 elements long and should be filled by a previous call to "sampleCoeffs". The returned matrix will be dimensioned (maxdeg + 1) x (maxdeg + 1). ◆ sampleCoeffCovariance() [2/2]
template<class Numeric >
Estimate the covariances of the coefficients returned by the "sampleCoeffs" function. The "coeffs" array should be at least maxdeg + 1 elements long and should be filled by a previous call to "sampleCoeffs" with the same location and scale. The returned matrix will be dimensioned (maxdeg + 1) x (maxdeg + 1). ◆ sampleCoeffs() [1/2]
template<class Numeric >
Build the coefficients of the orthogonal polynomial series for the given sample of points (empirical density function). The length of the array "coeffs" should be at least maxdeg + 1. Note that the coefficients are returned in the order of increasing degree (same order is used by the "series" function). ◆ sampleCoeffs() [2/2]
template<class Numeric >
Build the coefficients of the orthogonal polynomial series for the given sample of points (empirical density function). The length of the array "coeffs" should be at least maxdeg + 1. Note that the coefficients are returned in the order of increasing degree (same order is used by the "series" function). Before calculating the coefficients, the coordinates are shifted and scaled according to x_new = (x_original - location)/scale. The resulting coefficients are also divided by scale. ◆ sampleCoeffVars() [1/2]
template<class Numeric >
Estimate the variances of the coefficients returned by the "sampleCoeffs" function. The "coeffs" array should be at least maxdeg + 1 elements long and should be filled by a previous call to "sampleCoeffs". The "variances" array should have at least maxdeg + 1 elements. It will contain the respective variances upon return. ◆ sampleCoeffVars() [2/2]
template<class Numeric >
Estimate the variances of the coefficients returned by the "sampleCoeffs" function. The "coeffs" array should be at least maxdeg + 1 elements long and should be filled by a previous call to "sampleCoeffs" with the same location and scale. The "variances" array should have at least maxdeg + 1 elements. It will contain the respective variances upon return. ◆ sampleProductAverages()
template<class Numeric >
Unweighted averages of the pairwise products of the polynomial values for the given sample. The returned matrix will be symmetric and will have the dimensions (maxdeg + 1) x (maxdeg + 1). ◆ series()
Polynomial series ◆ twopoly()
Values of two orthonormal polynomials. Faster than calling "poly" two times. ◆ weight()
The weight function should be normalized Implemented in npstat::HermiteProbOrthoPoly1D, npstat::JacobiOrthoPoly1D, npstat::JohnsonOrthoPoly1D, npstat::DensityOrthoPoly1D, npstat::ChebyshevOrthoPoly2nd, npstat::ChebyshevOrthoPoly1st, npstat::ShiftedLegendreOrthoPoly1D, npstat::LegendreOrthoPoly1D, and npstat::ClassicalOrthoPoly1DFromWeight< Functor1D >. ◆ weightedPointsAverages()
template<class Pair , typename Real >
Averages of the polynomial values for the given sample of weighted points (not weighting by the polynomial weight function). The first element of the pair will be treated as the coordinate and the second element will be treated as weight. Weights must not be negative. The length of array "averages" should be at least maxdeg + 1. ◆ weightedSampleCoeffCovariance() [1/2]
template<class Pair >
Estimate the covariances of the coefficients returned by the "weightedSampleCoeffs" function. The "coeffs" array should be at least maxdeg + 1 elements long and should be filled by a previous call to "weightedSampleCoeffs". The returned matrix will be dimensioned (maxdeg + 1) x (maxdeg + 1). This code assumes that weights and coordinates are statistically independent from each other. ◆ weightedSampleCoeffCovariance() [2/2]
template<class Pair >
Estimate the covariances of the coefficients returned by the "weightedSampleCoeffs" function. The "coeffs" array should be at least maxdeg + 1 elements long and should be filled by a previous call to "weightedSampleCoeffs" with the same location and scale. The returned matrix will be dimensioned (maxdeg + 1) x (maxdeg + 1). This code assumes that weights and coordinates are statistically independent from each other. ◆ weightedSampleCoeffs() [1/2]
template<class Pair >
Build the coefficients of the orthogonal polynomial series for the given sample of weighted points (empirical density function). The first element of the pair will be treated as the coordinate and the second element will be treated as weight. Weights must not be negative. The length of the array "coeffs" should be at least maxdeg + 1. Note that the coefficients are returned in the order of increasing degree (same order is used by the "series" function). ◆ weightedSampleCoeffs() [2/2]
template<class Pair >
Build the coefficients of the orthogonal polynomial series for the given sample of weighted points (empirical density function). The first element of the pair will be treated as the coordinate and the second element will be treated as weight. Weights must not be negative. The length of the array "coeffs" should be at least maxdeg + 1. Note that the coefficients are returned in the order of increasing degree (same order is used by the "series" function). Before calculating the coefficients, the coordinates are shifted and scaled according to x_new = (x_original - location)/scale. The resulting coefficients are also divided by scale. ◆ weightedSampleCoeffVars() [1/2]
template<class Pair >
Estimate the variances of the coefficients returned by the "weightedSampleCoeffs" function. The "coeffs" array should be at least maxdeg + 1 elements long and should be filled by a previous call to "weightedSampleCoeffs". The "variances" array should have at least maxdeg + 1 elements. It will contain the respective variances upon return. This code assumes that weights and coordinates are statistically independent from each other. ◆ weightedSampleCoeffVars() [2/2]
template<class Pair >
Estimate the variances of the coefficients returned by the "weightedSampleCoeffs" function. The "coeffs" array should be at least maxdeg + 1 elements long and should be filled by a previous call to "weightedSampleCoeffs" with the same location and scale. The "variances" array should have at least maxdeg + 1 elements. It will contain the respective variances upon return. This code assumes that weights and coordinates are statistically independent from each other. ◆ xmin()
Support of the polynomial Implemented in npstat::JohnsonOrthoPoly1D, npstat::DensityOrthoPoly1D, npstat::StorablePolySeries1D, npstat::HermiteProbOrthoPoly1D, npstat::ChebyshevOrthoPoly2nd, npstat::ChebyshevOrthoPoly1st, npstat::JacobiOrthoPoly1D, npstat::ShiftedLegendreOrthoPoly1D, npstat::LegendreOrthoPoly1D, and npstat::ClassicalOrthoPoly1DFromWeight< Functor1D >. The documentation for this class was generated from the following file: Generated by |
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||